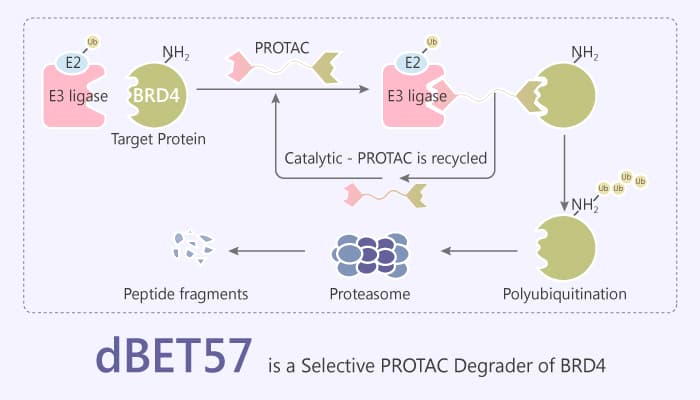
PROTACs (proteolysis-targeting chimeras) are bifunctional molecules that bring a target protein into spatial proximity with an E3 ubiquitin ligase to trigger target ubiquitination and subsequent proteasomal degradation. PROTAC recruits BRD4 to the E3 ubiquitin ligase cereblon leading to fast, efficient, and prolonged degradation of BRD4. As we all know, bromodomain protein BRD4 belongs to the bromodomain and extra terminal domain (BET) family of proteins. BRD4 contains two bromodomains (BD) at the N-terminus and an extra terminal domain (ET domain) at the C-terminus. Especially, BRD4 is a transcriptional repressor of autophagy and lysosomal function. Besides, BRD4 also plays a role in different biological processes, including memory formation, mitochondrial oxidative phosphorylation, and DNA damage response. dBET57 is a selective PROTAC BRD4 degrader.
PROTACs or heterobifunctional degrader molecules (referred to as degraders) typically comprise an E3 ligase binding scaffold. This often is an analog of thalidomide or a ligand to the von Hippel-Lindau tumor suppressor (VHL) protein. These proteins attach through a linker to another small molecule (hereafter target-moiety) that binds a target protein of interest. Recruitment of this target protein to the E3 ubiquitin ligase facilitates ubiquitination and subsequent degradation of the target protein. This principle successfully applies to several targets including the BET family (BRD2, BRD3, BRD4), RIPK2, BCR-ABL, FKBP12, BRD9, and ERRα. As a result, these effects above represents a promising new pharmacologic modality now widely explored in chemical biology and drug discovery.
dBET57 exhibits significant degradation of BRD4BD1 is inactive on BRD4BD2.
In particular, dBET57 is remarkably selective for BRD4BD1 in biochemical and cellular assays. Moreover, dBET57 exhibits potent effects on BRD4BD1 protein levels with DC50/5h of ~ 500 nM. Furthermore, dBET57 mediates recruitment to the CRL4CRBN E3 ubiquitin ligase.